Directions
The following steps describe how to calculate the optimal EQ filter and store it in syngo.MM Oncology. Siemens Healthineers provides the phantom scan and filter calculation service in some regions. For details please contact your local service representative.
1. Load the first attenuation corrected PET phantom dataset into syngo.MM Oncology.
2. Set the Units to Bq/ml from the bottom right corner menu of a segment displaying the phantom.

3. Using the wrench icon and selecting “Measurements”, select a threshold of 50% SUVmax.
4. Enable the RECIST/WHO + VOI Isocontour measurement tool.

5. Ensure that Max.eq is displayed in the measurement text:
a. Right click the RECIST/WHO + VOI Isocontour entry in the top right corner menu of a segment displaying the phantom.
b. Select VOI Isocontour Properties.
c. Check the Max and Max.eq options.
d. Uncheck all other options.

6. Use the VOI Isocontour tool to segment each hot sphere in the phantom. A large (~3cm) sphere can also be drawn in the background of the NEMA phantom (bottom left or bottom right, halfway between the air and the spheres) to provide background quality control for cross-calibration purposes.

7. Create a spreadsheet for EQ·PET optimization, for example the figure below. The spreadsheet should input the known hot sphere activity concentration at the beginning of the scan, the EQ filter size, the target recovery curve (target scanner or EANM reference standard), and the measured Max.eq values for the respective spheres. The difference between the target recovery curve and the recovery curve measured in syngo.MM Oncology with a specific EQ filter size should be calculated as the mean absolute percentage difference. For the latest EANM reference recovery coefficients refer to the EANM guidelines.

8. If the EQ·PET calibration is being performed to match another PET/CT system (scanner B) instead of the EARL parameters, then the measured Max.eq values for the respective spheres from a similar acquisition on scanner B should replace those in the “Target (kBq/ml)” column in the spreadsheet.
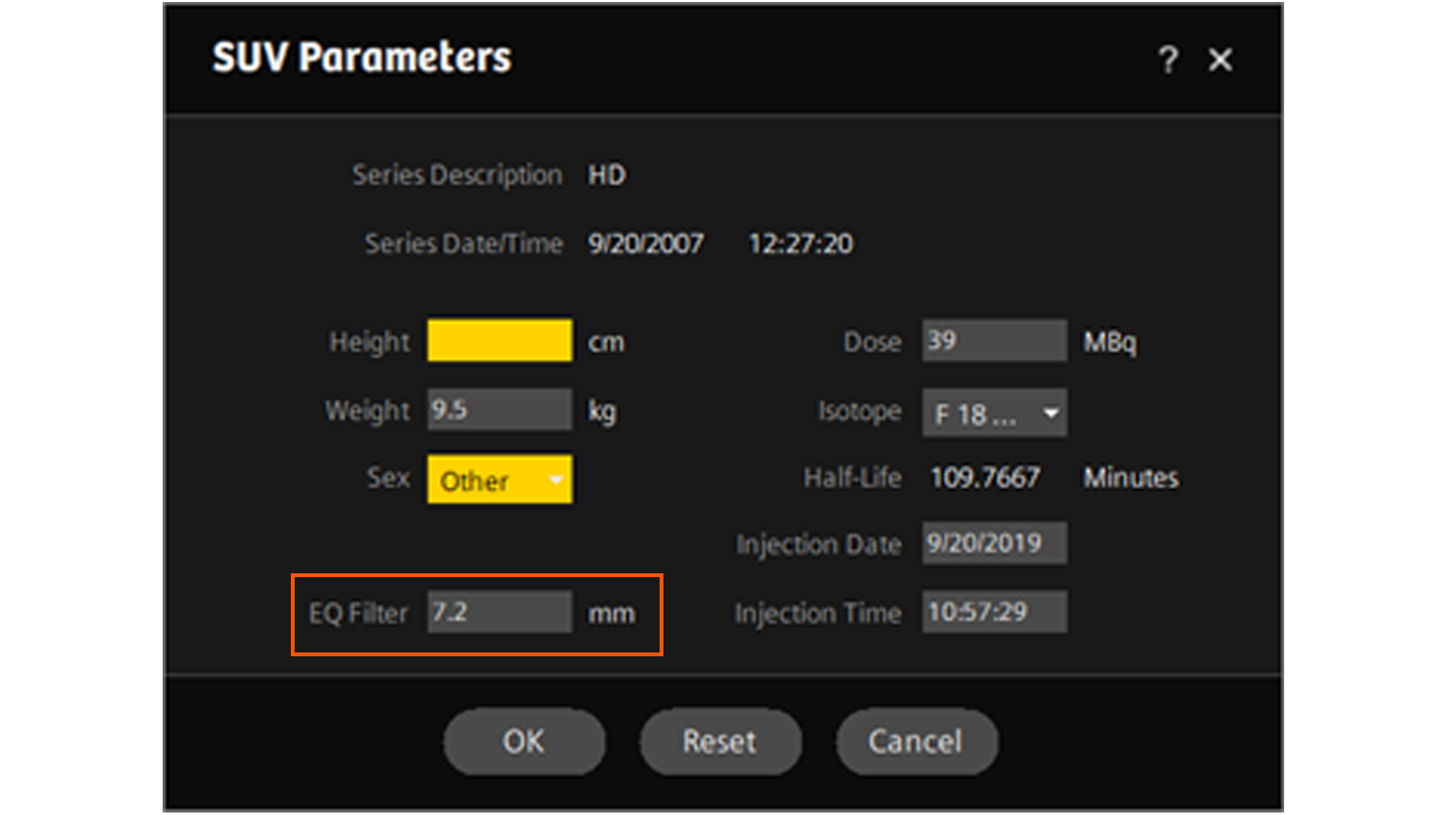
9. Return to syngo.MM Oncology and increase the EQ filter setting by 0.1 mm. The EQ filter is available by left clicking on the SUV Parameters option on the bottom right corner menu in the segment displaying the phantom.


10. Measure the Max.eq in the NEMA phantom spheres.

11. Enter the Max.eq values for the six spheres in the next open spreadsheet column, with the EQ filter entered at the top of the column for reference. Observe the mean absolute % difference with the EARL target value under the column.
12. Continue increasing the EQ filter in 0.1 mm increments in syngo.MM Oncology, entering Max.eq values and recording the EQ filter value after each increment in a new column on the spreadsheet. Note that the Max.eq values may only start differing from the first (0 mm) column after several increments of 0.1 mm. In practice there is typically one minimum error, so initial increments can be larger than 0.1 mm.

13. When the mean absolute % difference with EARL target value starts to increase, you can stop incrementing the EQ filter. To be sure that the minimum % difference is captured, check that the % difference with EARL target increases continuously at +-0.1 mm and +-0.2 mm from minimum error.

14. Record the EQ filter number which gave you the lowest % difference value.
15. An EQ·PET filter should be computed for each reconstruction protocol used at the institution.
16. Repeat these steps for the other phantom acquisitions if multiple scans were performed. Note that the EARL recommendation is three acquisitions. There should not be a large variation in the EQ filter values for the different acquisitions with identical reconstruction protocols (1 – 2 mm difference at most).
17. Calculate the mean of the three EQ filter values and record this number, noting the reconstruction protocol that was used. Enter this final filter value into the EQ filter field in syngo.MM Oncology to ensure that it is stored. The EQ filter is assigned uniquely based on scanner model, reconstruction method, convolution kernel, rows, columns, pixel spacing and slice thickness.